ELIXIR Interoperability Platform: A Foundation for Integrated Life Science Data Management
March 26, 2026
Life science research is increasingly based on large, diverse, and geographically distributed datasets. To use such data effectively, it is not enough to store it or provide access - it is also necessary to be able to link, interpret, and reuse it across different systems. To address these challenges, the European life science data infrastructure has been established, bringing together national research infrastructures into a unified ecosystem.
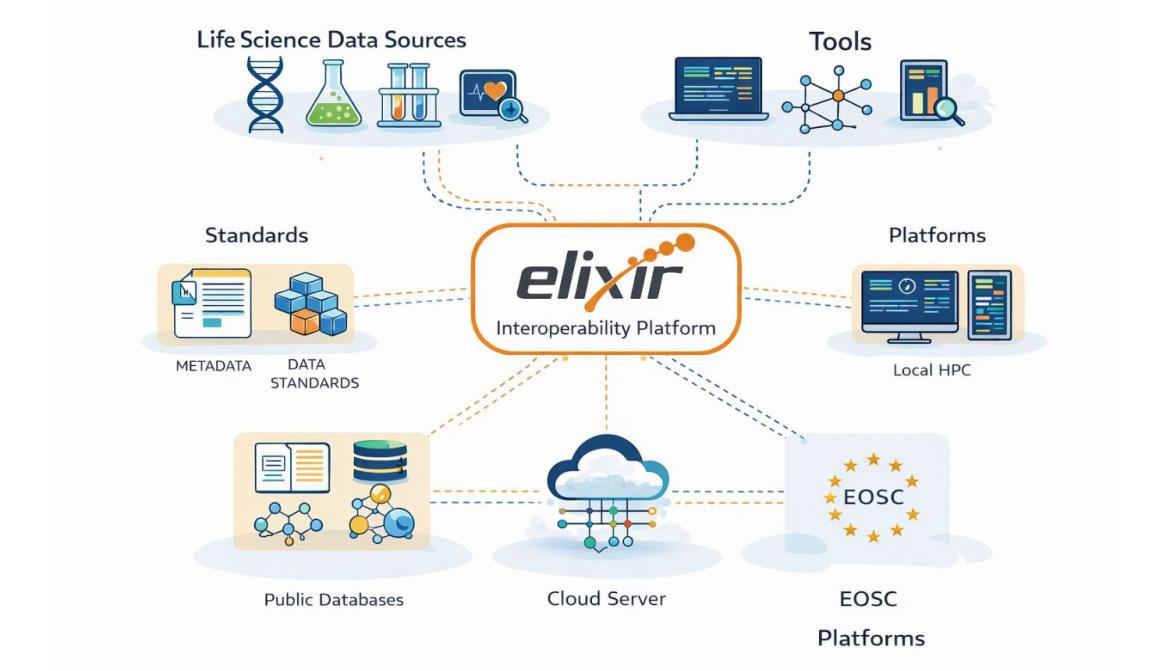
ELIXIR operates through five interconnected platforms: Data, Tools, Compute, Interoperability, and Training. This article focuses on the Interoperability Platform, whose main objective is to ensure that life science data and digital resources can work together regardless of their origin, format, or the infrastructure used.

Data Interoperability as a Foundation for Modern Science
The ELIXIR Interoperability Platform focuses on standards, metadata, ontologies, and data management principles that enable the integration of diverse systems. This is particularly important in bioinformatics, where researchers often use data from different laboratories, databases, and countries. The Interoperability Platform helps both humans and machines to find, access, integrate, and analyse life science data. It encourages the life sciences community to adopt internationally standardised file formats, metadata, terminologies (vocabularies), and identifiers, thereby enabling analytical tools to be linked with primary data.
The platform’s recommended resources, for example, on data linking, metadata usage, and the development of interoperable data collections, are available here.
Examples of interoperability can be viewed here.
The platform promotes the implementation of the FAIR data principles, ensuring that data are Findable, Accessible, Interoperable, and Reusable. These principles have become one of the central elements of modern scientific data management.
The activities of the Interoperability Platform are based on several key areas, which together form a unified ecosystem for data integration.
The platform promotes the use of common standards for life science data - such as in genomics, proteomics, or metabolomics - by applying standardised terminology, consistent data descriptions, standardised metadata, and shared data formats. Ontologies provide a unified terminological framework that allows different datasets to retain clear semantic meaning. For example, biological processes, diseases, or molecular mechanisms are described using standardised concepts. This not only facilitates data integration but also improves automated data analysis and enables more effective machine learning approaches.
The platform also develops guidelines for research data management. This includes the preparation of data management plans, ensuring data quality, documenting data, and publishing data in open infrastructures. These activities help researchers organise their data so that it remains usable in the future.
The ELIXIR Interoperability Platform works closely with other ELIXIR platforms, such as the Data Platform to ensure that data resources and metadata are properly described; the Tools Platform so that analytical tools can directly use standardised data; and the Compute Platform so that infrastructures can process data across different computing environments. This integration creates a unified European bioinformatics infrastructure in which data, tools, and computational resources operate as an interconnected ecosystem.
The Interoperability Platform also actively collaborates with international initiatives that develop global data standards. This is particularly important, as life science data are often used in international research. The platform’s activities are also closely linked to the European Open Science Cloud (EOSC) initiative, which aims to create a federated infrastructure for the storage, sharing, and analysis of scientific data. Interoperability principles are essential for such an infrastructure to function effectively, enabling data exchange across different scientific disciplines and institutions.
The ELIXIR Interoperability Platform forms the foundation for modern life science research by ensuring that data do not remain isolated within individual systems, but can be connected, analysed, and interpreted within a broader scientific context. As a result, researchers can combine data from different experiments, use shared analytical workflows, enable reproducible research, and accelerate discoveries in the life sciences. Interoperability is becoming one of the most essential elements of digital science, as it ensures that scientific data becomes a long-term, usable, and integrable resource.
Although Latvia is not yet a full member of ELIXIR, its research institutions can already participate in several platform-related activities. This is possible through involvement in European research projects, collaborative initiatives, or the development of data infrastructures. Latvia’s participation is particularly significant in projects related to genomics data infrastructures and the European Open Science Cloud, where ELIXIR is one of the key partners.
On June 1, 2025, Riga Stradiņš University (RSU) launched the project “RSU Participation in the Horizon Europe Programme” (Project No. 1.1.1.5/3/25/I/014), which also aims to promote Latvia’s integration into the ELIXIR infrastructure. This initiative may open up opportunities for Latvian researchers to participate more actively in the development of the European life sciences data infrastructure. Full membership of Latvia in ELIXIR in the future would allow national research infrastructures to directly engage in the work of the Interoperability Platform, develop common data standards, and integrate Latvia’s digital research infrastructure into the European bioinformatics ecosystem.
